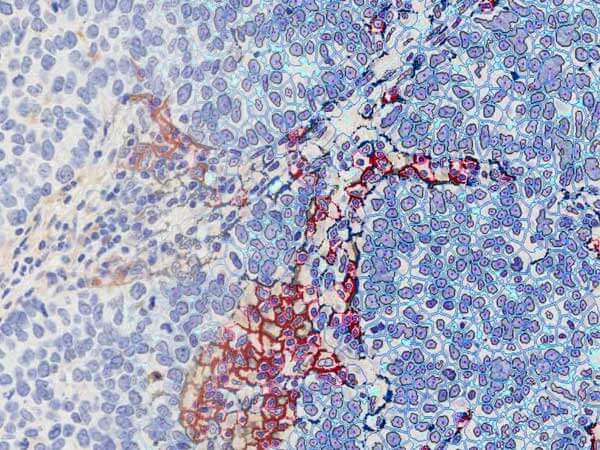
Quantifying PD-L1 Spatial Distribution Signatures for Patient Selection Approaches
Introduction
- Inhibitors of inflammatory checkpoints, such as PD-L1 inhibitors, have demonstrated great promise in preclinical and clinical studies. This therapeutic paradigm focuses on controlling natural inflammatory checkpoints to stimulate an elevated inflammatory response against the tumor by increasing anti-tumor inflammatory cell infiltrates in the tumor microenvironment (TME) or decreasing inflammatory suppressor infiltrates.
- The proteins which control these processes can be found in the tumor cells, cells in the TME, or in both locales. Positive cells are often assessed in a qualitative or semi-quantitative manner using immunohistochemistry and evaluation of a limited number of representative microscopy fields across a particular tissue compartment (tumor vs stroma) or the whole tissue area.
- However, a more specific locale measurement of the inflammatory suppressors, such as PD-L1, may be more revealing than estimating the tumor-wide dispersion of an inflammatory cell type. Unfortunately, the intricate spatial relationships and the often complex distribution of inflammatory cells in tissues pose significant challenges for a meaningful evaluation.
- We have developed an approach which can quantify these spatial relationships for a biologically meaningful score.