Abstract:
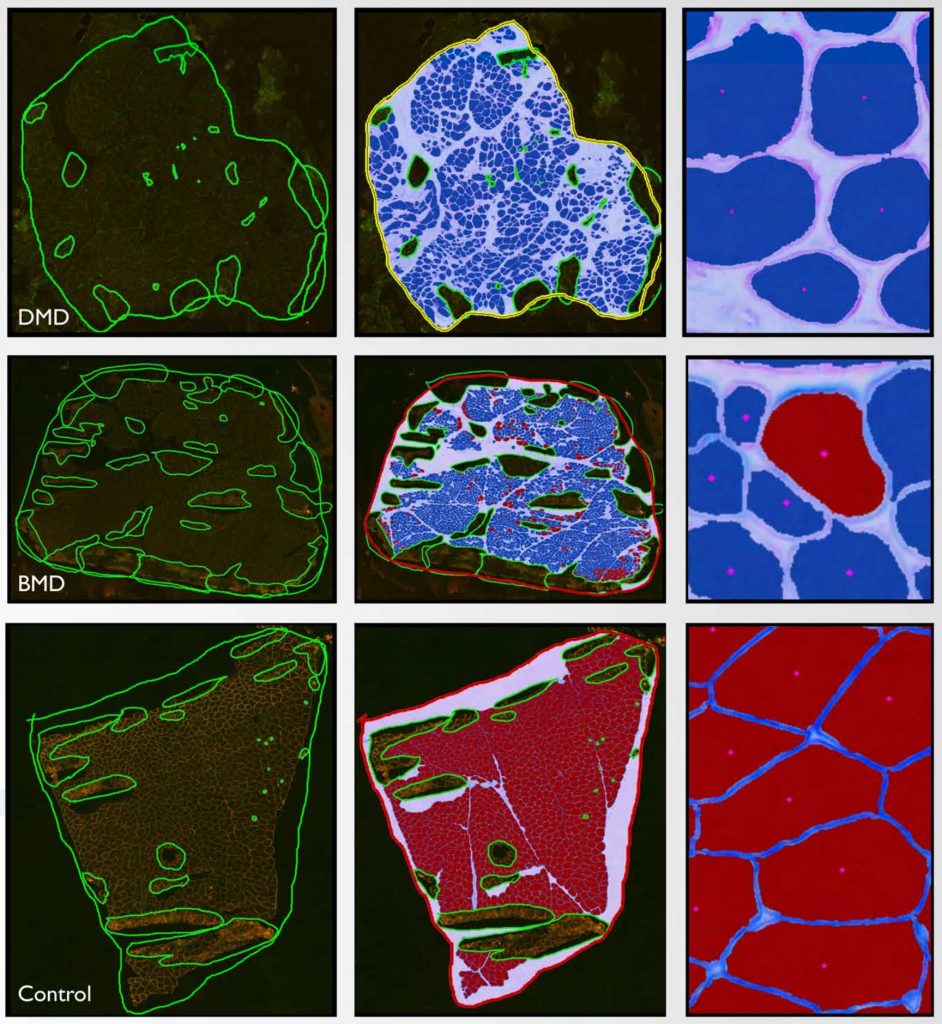
Quantification of skeletal muscle fiber parameters is challenging, but important for the evaluation of a variety of neuromuscular diseases. Various endpoints from individual fiber characteristics to whole tissue determinations may be required to better understand disease biology and/or treatment efficacy. In the context of muscular dystrophy studies, assessment of dystrophin expression in muscle fibers has largely been qualitative or semi-quantitative assessments of representative areas of an entire tissue section. To provide more robust quantitation of muscle fiber endpoints, an automated method (MuscleMap™) was created to quantify a number of group and individual muscle fiber parameters. To evaluate algorithm efficacy, MuscleMap was applied to a test cohort of dystrophic muscle biopsies. Duchenne and Becker muscular dystrophy (DMD and BMD, respectively) are rare genetic diseases that result from mutations in the DMD gene, which encodes the dystrophin protein, leading to progressive muscle degeneration, muscle weakness and fatigue, and premature death. Muscle biopsy cryosections derived from DMD and BMD patients and from healthy control individuals were assessed using the MuscleMap algorithm. Numerous parameters relating to staining intensity, membrane staining completeness, and morphometric presentation of dystrophin in individual muscle fibers were quantified in dual label, immunofluorescence-stained sections. A number of parameters, including mean dystrophin staining intensity and dystrophin membrane staining completeness, were significantly different in DMD and BMD tissue when compared to normal controls, and are promising biomarkers for understanding biology of the disease. From early drug development to clinical trials, automated quantification of dystrophin expression in muscle fibers is an attractive endpoint given the mechanism of action for current promising therapies.